Large-Scale proteome-wide mendelian randomization identifies novel proteins for glaucoma and related traits
关键词
摘要
全文
INTRODUCTION
METHODS
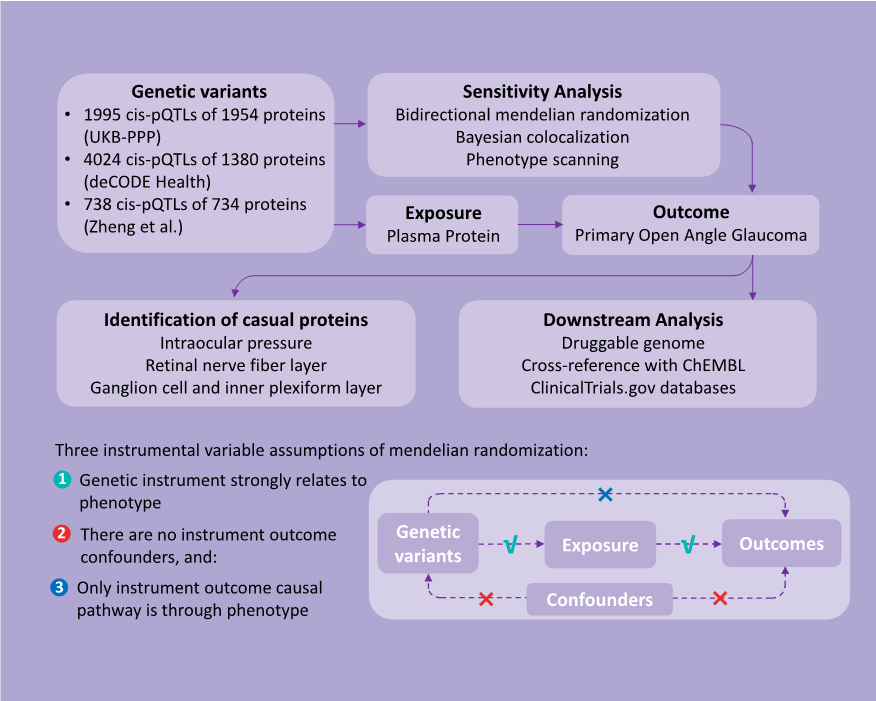
Study Design
Data Sources
STATISTICAL ANALYSIS
Two-sample MR Analysis
Reverse Causality Detection
Bayesian Colocalization Analysis
We conducted colocalization analysis to discern the impact of horizontal pleiotropy from genetic IVs on the identified causal links.[26] For this analysis, we utilized a Bayesian framework to evaluate posterior probabilities for five scenarios regarding the genetic overlap between plasma proteins and POAG. The scenarios ranged from no shared causal variants (H0) to specific causal variants for proteins (H1) or POAG (H2), independent causal variants affecting both (H3), and a common causal variant for both (H4). A posterior probability of (PH4) ≥ 0.8 indicates strong colocalization, whereas 0.8 > PH4 ≥ 0.5 suggestes moderate support. This was performed using the “coloc” package in R (version 4.4.1).Phenotype Scanning
Utilizing the “phenoscanner” R package, we explored large-scale genetic association studies for links between our identified pQTLs and POAG-related traits.[27] SNPs that showed pleiotropy met stringentcriteria: genome-wide significance (P < 5×10-8), association studies in European populations, and links to established POAG risk factors, including metabolic traits, proteins, or clinical features.Identification in POAG-related Traits
Downstream Analysis of Drug Target Proteins
Data Availability
RESULTS
Mendelian Randomization Between Plasma Proteins and POAG
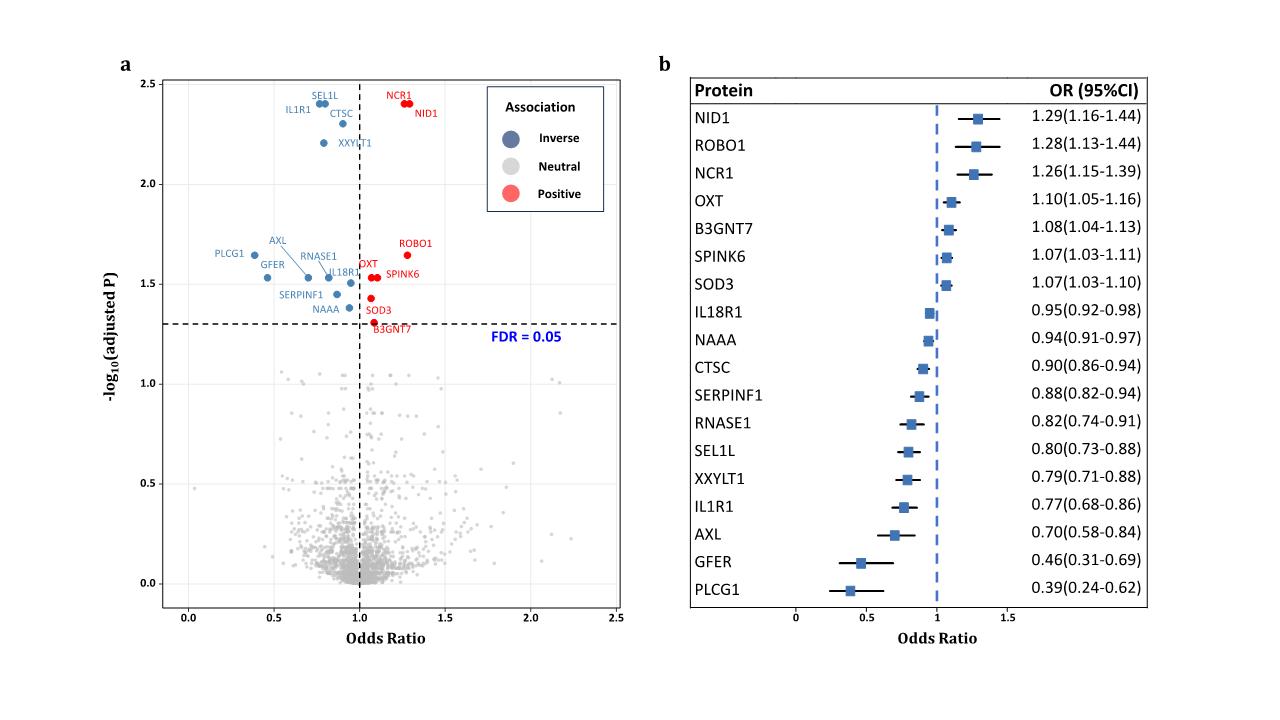
Table 1 Plasma proteins significantly associated with primary open-angle glaucoma after false discovery rate (FDR) correction
|
Protein |
Chr |
Position |
rs number cis-pQTL |
EAF |
EA |
MR analysis |
Source
|
||||||
|
β (SE) |
OR |
95%CI |
P value |
R2 |
F statistics |
|
|||||||
|
AXL |
19 |
41233275 |
rs66841352 |
0.40 |
C |
-0.36(0.09) |
0.70 |
0.58-0.84 |
1.62×10-4 |
0.014 |
473.03 |
UKB-PPP |
|
|
B3GNT7 |
2 |
231398416 |
rs2290130 |
0.23 |
A |
0.08(0.02) |
1.08 |
1.04-1.13 |
3.77×10-4 |
0.169 |
6888.59 |
UKB-PPP |
|
|
GFER |
16 |
1973321 |
rs61516948 |
0.13 |
T |
-0.77(0.20) |
0.46 |
0.31-0.69 |
1.50×10-4 |
0.003 |
89.38 |
UKB-PPP |
|
|
PLCG1 |
20 |
41168825 |
rs753381 |
0.47 |
C |
-0.95(0.24) |
0.39 |
0.24-0.62 |
7.70×10-5 |
0.002 |
70.03 |
deCODE |
|
|
SEL1Lb |
14 |
81506097 |
rs11499034 |
0.01 |
C |
-0.22(0.05) |
0.80 |
0.73-0.88 |
5.05×10-6 |
0.044 |
1538.02 |
UKB-PPP |
|
|
SERPINF1 |
17 |
1666253 |
rs62088172 |
0.35 |
T |
-0.13(0.04) |
0.88 |
0.82-0.94 |
2.54×10-4 |
0.088 |
316.65 |
Sun |
|
|
CTSCa, b |
11 |
88337746 |
rs11600158 |
0.10 |
G |
-0.10(0.02) |
0.90 |
0.86-0.94 |
1.06×10-5 |
0.146 |
5787.36 |
UKB-PPP |
|
|
88066714 |
rs55897509 |
0.92 |
A |
0.085 |
509.91 |
Emilsson |
|||||||
|
IL18R1a |
2 |
102377596 |
rs10190555 |
0.77 |
G |
-0.05(0.01) |
0.95 |
0.92-0.98 |
1.86×10-4 |
0.228 |
9955.02 |
UKB-PPP |
|
|
101625575 |
rs115232861 |
0.03 |
C |
0.006 |
226.86 |
deCODE |
|||||||
|
103035044 |
rs1420106 |
0.78 |
G |
0.270 |
1221.57 |
Sun |
|||||||
|
102877724 |
rs183611009 |
0.02 |
G |
0.006 |
227.53 |
deCODE |
|||||||
|
102030778 |
rs187572594 |
0.02 |
G |
0.002 |
63.20 |
deCODE |
|||||||
|
101829431 |
rs55715763 |
0.06 |
G |
0.004 |
133.11 |
deCODE |
|||||||
|
102264346 |
rs55871806 |
0.15 |
C |
0.088 |
3399.96 |
deCODE |
|||||||
|
102549866 |
rs72995641 |
0.21 |
A |
0.031 |
1126.2 |
deCODE |
|||||||
|
101525580 |
rs75094400 |
0.02 |
C |
0.003 |
101.34 |
deCODE |
|||||||
|
102971459 |
rs80339564 |
0.03 |
C |
0.003 |
116.82 |
deCODE |
|||||||
|
IL1R1a, b |
2 |
102156623 |
rs3917238 |
0.30 |
T |
-0.27(0.06) |
0.77 |
0.68-0.86 |
4.22×10-6 |
0.015 |
528.97 |
UKB-PPP |
|
|
102138349 |
rs6706048 |
0.28 |
C |
0.012 |
440.56 |
deCODE |
|||||||
|
102746276 |
rs7588201 |
0.72 |
A |
0.007 |
39.29 |
Emilsson |
|||||||
|
NAAAa |
4 |
75932965 |
rs112197434 |
0.23 |
T |
-0.06(0.02) |
0.94 |
0.91-0.97 |
3.00×10-4 |
0.141 |
5525.12 |
UKB-PPP |
|
|
76102346 |
rs12331871 |
0.17 |
G |
0.016 |
568.16 |
deCODE |
|||||||
|
75906793 |
rs1394919 |
0.29 |
A |
0.148 |
6148.47 |
deCODE |
|||||||
|
76209167 |
rs76229059 |
0.17 |
G |
0.005 |
190.32 |
deCODE |
|||||||
|
76848231 |
rs9996608 |
0.30 |
T |
0.119 |
445.59 |
Sun |
|||||||
|
NCR1a, b |
19 |
55419632 |
rs143981324 |
0.90 |
T |
0.23(0.05) |
1.26 |
1.15-1.39 |
1.99×10-6 |
0.013 |
74.26 |
Emilsson |
|
|
54904812 |
rs7255591 |
0.09 |
C |
0.031 |
1073.76 |
UKB-PPP |
|||||||
|
NID1a, b |
1 |
236099073 |
rs17554536 |
0.18 |
A |
0.26(0.06) |
1.29 |
1.16-1.44 |
6.72×10-6 |
0.002 |
67.92 |
deCODE |
|
|
236104574 |
rs58074293 |
0.02 |
A |
0.011 |
379.94 |
deCODE |
|||||||
|
236067017 |
rs76183323 |
0.01 |
A |
0.012 |
402.08 |
UKB-PPP |
|||||||
|
OXTa |
20 |
3067658 |
rs6115776 |
0.40 |
C |
0.10(0.03) |
1.10 |
1.05-1.16 |
1.61×10-4 |
0.059 |
2208.44 |
deCODE |
|
|
3100953 |
rs857244 |
0.46 |
C |
0.001 |
41.77 |
deCODE |
|||||||
|
3049890 |
rs877172 |
0.66 |
T |
0.125 |
776.73 |
Emilsson |
|||||||
|
RNASE1a |
14 |
20789957 |
rs11624082 |
0.26 |
T |
-0.20(0.05) |
0.82 |
0.74-0.91 |
1.16×10-4 |
0.001 |
50.25 |
deCODE |
|
|
20814883 |
rs12897030 |
0.30 |
T |
0.016 |
592.52 |
deCODE |
|||||||
|
21280678 |
rs17254387 |
0.69 |
A |
0.022 |
72.52 |
Sun |
|||||||
|
20828671 |
rs188152481 |
0.02 |
C |
0.002 |
63.19 |
deCODE |
|||||||
|
20722147 |
rs2002078 |
0.41 |
T |
0.004 |
158.96 |
deCODE |
|||||||
|
20627658 |
rs75124405 |
0.01 |
A |
0.002 |
66.29 |
deCODE |
|||||||
|
ROBO1a |
3 |
78946828 |
rs331172 |
0.49 |
G |
0.25(0.06) |
1.28 |
1.13-1.44 |
6.78×10-5 |
0.003 |
88.58 |
deCODE |
|
|
78725134 |
rs3773233 |
0.21 |
T |
0.014 |
485.44 |
deCODE |
|||||||
|
78735620 |
rs3773244 |
0.20 |
A |
0.020 |
687.56 |
UKB-PPP |
|||||||
|
79787885 |
rs62256944 |
0.10 |
A |
0.002 |
58.34 |
deCODE |
|||||||
|
77848463 |
rs73103884 |
0.02 |
G |
0.001 |
45.09 |
deCODE |
|||||||
|
78741405 |
rs73111654 |
0.01 |
A |
0.002 |
81.48 |
deCODE |
|||||||
|
79369917 |
rs7639769 |
0.46 |
T |
0.003 |
100.95 |
deCODE |
|||||||
|
SOD3a |
4 |
24800212 |
rs1799895 |
0.01 |
G |
0.06(0.02) |
1.07 |
1.03-1.10 |
2.42×10-4 |
0.133 |
5170.20 |
UKB-PPP |
|
|
24802616 |
rs2695234 |
0.08 |
G |
0.031 |
1145.21 |
deCODE |
|||||||
|
25235769 |
rs313548 |
0.30 |
A |
0.003 |
109.25 |
deCODE |
|||||||
|
25664005 |
rs76930256 |
0.02 |
T |
0.005 |
190.96 |
deCODE |
|||||||
|
24444429 |
rs77514718 |
0.01 |
C |
0.014 |
505.01 |
deCODE |
|||||||
|
24639073 |
rs80234081 |
0.03 |
C |
0.021 |
751.96 |
deCODE |
|||||||
|
SPINK6a |
5 |
148263754 |
rs116664727 |
0.13 |
A |
0.07(0.02) |
1.07 |
1.03-1.11 |
1.41×10-4 |
0.006 |
223.76 |
deCODE |
|
|
148186561 |
rs12717962 |
0.55 |
G |
0.072 |
2630.07 |
UKB-PPP |
|||||||
|
147294723 |
rs137864680 |
0.02 |
A |
0.001 |
50.58 |
deCODE |
|||||||
|
148310721 |
rs139120114 |
0.01 |
C |
0.014 |
511.80 |
deCODE |
|||||||
|
147603178 |
rs1432688 |
0.92 |
G |
0.155 |
605.02 |
Sun |
|||||||
|
149047717 |
rs148402978 |
0.02 |
T |
0.003 |
113.83 |
deCODE |
|||||||
|
147734214 |
rs148632877 |
0.01 |
T |
0.002 |
70.11 |
deCODE |
|||||||
|
148183699 |
rs74765295 |
0.01 |
T |
0.007 |
248.34 |
deCODE |
|||||||
|
147882756 |
rs75666474 |
0.05 |
A |
0.003 |
93.34 |
deCODE |
|||||||
|
148453118 |
rs75666559 |
0.04 |
T |
0.007 |
259.13 |
deCODE |
|||||||
|
148057440 |
rs7719473 |
0.03 |
A |
0.057 |
2144.61 |
deCODE |
|||||||
|
XXYLT1a, b |
3 |
195287123 |
rs1538767 |
0.46 |
C |
-0.23(0.05) |
0.79 |
0.71-0.88 |
1.58×10-5 |
0.016 |
564.92 |
deCODE |
|
|
195619664 |
rs534994266 |
0.09 |
A |
0.002 |
80.40 |
deCODE |
|||||||
|
194783033 |
rs55947051 |
0.84 |
T |
0.010 |
56.46 |
Emilsson |
|||||||
|
195072715 |
rs7635512 |
0.16 |
C |
0.007 |
240.77 |
deCODE |
|||||||
Source indicates the protein GWAS providing the estimate of the effect of the cis-pQTL on protein level. Results express changes in primary open angle glaucoma risk per 1-SD increase in protein level. EA=effect allele; EAF=effect allele frequency; β=effect on diabetic retinopathy; OR=odds ratio.
a Mendelian randomization using the inverse variance weighted methods.
b Plasma protein passed Bonferroni correction (P < 2.12×10-5).
Multiple Sensitivity Analyses
Table 2. Summary of reverse causality detection, Bayesian co-localization analysis and phenotype scanning on 18 potential causal proteins.
|
Protein |
Bidrectional MR |
Co-localization |
Previously reported associations |
||
|
OR(95%CI) |
P value |
PPH3 |
PPH4 |
||
|
SEL1L |
1.00(0.98-1.02) |
0.80 |
0.002 |
0.995 |
lipid metabolism |
|
ROBO1 |
1.00(0.98-1.03) |
0.92 |
0.065 |
0.865 |
NA |
|
AXL |
1.00(0.99-1.00) |
0.12 |
0.052 |
0.858 |
NA |
|
NID1 |
1.00(0.97-1.02) |
0.73 |
0.039 |
0.806 |
NA |
|
GFER |
1.01(0.99-1.03) |
0.34 |
0.125 |
0.804 |
NA |
|
NCR1 |
1.00(0.97-1.02) |
0.81 |
0.335 |
0.505 |
NA |
|
SERPINF1 |
0.96(0.91-1.02) |
0.20 |
0.366 |
0.476 |
Body measurement |
|
CTSC |
0.99(0.97-1.01) |
0.36 |
0.323 |
0.431 |
NA |
|
SPINK6 |
1.00(0.98-1.02) |
0.94 |
0.399 |
0.260 |
NA |
|
B3GNT7 |
1.01(0.99-1.04) |
0.25 |
0.322 |
0.210 |
Body measurement |
|
RNASE1 |
0.97(0.93-1.02) |
0.29 |
0.117 |
0.193 |
Cervix uteri disease |
|
NAAA |
0.99(0.97-1.01) |
0.36 |
0.181 |
0.116 |
NA |
|
IL1R1 |
1.00(0.97-1.02) |
0.75 |
0.354 |
0.076 |
Blood cell, asthma, hayfever, rhinitis, eczema, bronchitis, emphysema |
|
IL18R1 |
1.02(1.00-1.03) |
0.07 |
0.340 |
0.026 |
Blood cell, inflammatory bowel disease |
|
PLCG1 |
1.01(0.96-1.06) |
0.68 |
0.092 |
0.025 |
Blood cell, lipid metabolism |
|
XXYLT1 |
0.96(0.91-1.02) |
0.20 |
0.520 |
0.022 |
NA |
|
SOD3 |
0.99(0.97-1.01) |
0.42 |
0.506 |
0.009 |
Vascular dementia |
|
OXT |
1.02(0.96-1.08) |
0.54 |
0.018 |
0.004 |
NA |
OR=odds ratio; NA=not applicable.
Identification of Plasma Proteins in IOP, RNFL, and GCIPL
The Druggability and Development Status of POAG-causal Protein
Table 3. Summary of druggability and clinical development activity for primary open angle glaucoma associated with causal associations on MR analysis.
|
Target |
Compound name |
Clinical development activities |
|
Approved |
||
|
AXL |
GILTERITINIB (CHEMBL3301622)b |
Approved for neoplasms, acute myeloid leukemia, myeloid leukemia; Phase II: myelodysplastic syndromes, blast crisis; Phase I: non-smal-cell lung carcinoma, kidney diseases, liver diseases |
|
IL1R1 |
ANAKINRA (CHEMBL1201570)b |
Approved: immune system diseases; cryopyrin-associated periodic syndromes, rheumatoid arthritis; Phase III: metabolic syndrome, giant cell arteritis, pneumonia, severe acute respiratory syndrome, myocardial infarction, mucocutaneous lymph node syndrome, familial mediterranean fever, influenza, dermatitis, allergic contact; Phase II: brain injuries, lymphoma, follicular, multiple myeloma, pericarditis, diabetes mellitus, Still's disease, uveitis, hvidradenitis suppurativa, myositis, alcoholic hepatitis, macrophage activation syndrome, gout, dermatomyositis, sarcoidosis, amyotrophic lateral sclerosis, infections, hypoglycemia, type 1 diabetes mellitus, osteoarthritis, juvenile arthritis, heart failure, cerebral hemorrhage, polycystic ovary syndrome, myocarditis, labyrinth diseases, chronic fatigue syndrome, chronic renal insufficiency; Phase I: castration-resistant prostatic neoplasms, pancreatic neoplasms, knee injuries, blepharitis, HIV infections, multiple sclerosis, breast neoplasms, pulmonary hypertension, pain, smoldering multiple, myeloma, asthma, rectal neoplasms, chronic B-cell leukemia lymphocytic, hearing loss, neoplasms |
|
In development |
||
|
AXL |
BEMCENTINIB (CHEMBL3809489)b |
Phase II: adenocacinoma of lung; mesothelioma; severe acute respiratory; inflammatory breast neoplasms; Phase I: myelodysplastic syndromes; acute leukemia myeloid; pancreatic neoplasms; non-small-cell lung carcinoma; melanoma |
|
AXL |
ENAPOTAMAB VEDOTIN (CHEMBL4297987)b |
Phase I: non-small-cell lung carcinoma |
|
AXL |
BPI-9016 (CHEMBL3545236)b |
Phase I: non-small-cell lung carcinoma; neoplasms |
|
CTSC |
BRENSOCATIB (CHEMBL3900409)b |
Phase III: cystic fibrosis; COVID-19; Phase II: bronchiectasis |
|
IL18R1 |
IBOCTADEKIN (CHEMBL2108034)b |
Phase II: melanoma; Phase I: neoplasms; lymphoma non-hodgin; lymphoma |
|
IL1R1 |
MEDI-8968 (CHEMBL2109607)b |
Phase II: hidradenitis suppurativa; chronic obstructive pulmonary disease |
|
IL1R1 |
AMG-108 (CHEMBL2109458)b |
Phase II: rheumatoid arthritis; diabetes mellitus; osteoarthritis |
|
OXT |
OXYTOCINc |
Phase III: Prader-Willi syndrome (NCT02804373), socially adaptive mirroring (NCT03640156); Phase II: central diabetes insipidus (NCT06036004), schizophrenia (NCT01699997) |
|
Druggablea |
||
|
AXL, GFER, SERPINF1, CTSC, IL18R1, IL1R1, NAAA, NCR1, NID1, OXT, RNASE1, SOD3, SPINK6 |
||
|
Not currently listed as druggablea |
||
|
B3GNT7, PLCG1, SEL1L, ROBO1, XXYLT1 |
||
a Data from druggable gene list.
b Data from ChEMBL release 33 (compound ID in brackets).
c Data from ClinicalTrials.gov.
DISCUSSION
Summary of Findings
MR-derived novel biomarkers and potential targets for POAG
Nidogen-1 (NID1), a key basement membrane glycoprotein, has been found to be as positively associated with POAG risk.[39-40] Increased nidogen presence in glaucoma patients suggests its involvement in extracellular matrix dynamics and aqueous efflux obstruction.[41] The specific mechanisms by which NID1 contributes to POAG remain to be clarified, thereby highlighting a crucial area for future research. Our study found that higher levels of GFER, which is a protein vital for mitochondrial integrity, is correlated with reduced POAG risk.[42-44] This suggests GFER’s protective role in preventing oxidative stress and mitochondrial dysfunction, both of which contributes in glaucoma progression.[44-45] The potential of GFER as a therapeutic target for POAG highlights the need for additional research into its mechanisms of action.